Bjorn’s interdisciplinary background ranges from computer science to visualisation and design.
Bjorn received his BSc in Media Informatics and Design in 2004 from Bielefeld University and Bielefeld University of Applied Sciences (Gestaltung). There he also completed an MA Interdisciplinary Media Sciences in 2006 and initiated the CELLmicrocosmos project 'Starting from an animated cell model'.
From 2009 to 2015 he worked and taught as a research assistant in the Bio-/Medical Informatics Department at Bielefeld University while creating the lecture module 'Interdisciplinary Cell Visualisation and Modelling' and finishing his PhD thesis on Integrative Cell Modelling.
In 2015, he started as a Research Fellow at Monash University in Melbourne and then at the University of Konstanz.
Gallery
More information
Research interests
Bjorn’s research focuses on the combination of information and scientific visualisation, driven by the exploration of abstract data and associated structures to increase the understanding of underlying processes. His major focus in recent years was life science related data, such as visualisation and simulation of structure and dynamics of cell membranes, movement data of animals, localisation data of proteins or abstract posture representations.
Part of his educational and research work includes the development and supervision of software frameworks which provide interactive modelling and exploration capabilities.
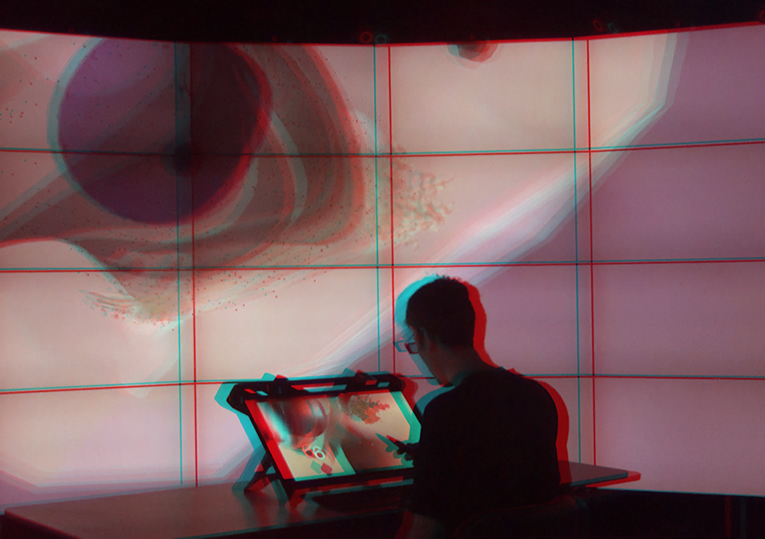
An important factor of most of these approaches is the support of stereoscopic visualisation/virtual reality-related exploration which comes with challenges on the software as well as hardware side. For this purpose, Bjorn has set up stereoscopic display systems on different scales – from single to multi-display environments.
Research funding
DAAD Grant – Project Exchange Program: 'Deciphering polymyxin resistance in Pseudomonas aeruginosa', with Y. Zhu, J. Li and F. Schreiber, 2018–19.
Microbiology@Monash Collaborative Grant, Monash University, Melbourne, with Y. Zhu, 2016.
ERASMUS PLUS Stipend, Research Stay at Verona University, 2015.
NSFC Award, Research Fund for International Young Scientists, Hangzhou, Zhejiang University, with M. Chen, 2014–15.
European Science Foundation (ESF) Fellowships for the EMBO Workshop VizBi, 2010.
DFG Stipend in the DFG Graduate College Bioinformatics Bielefeld (GK635), 2007–9.
Awards
Best Animation of Show at Stereoscopic Displays and Applications XXVII., San Francisco: 'Chlamydomonas reinhardtii – From Biological Cells to Biofuel', with N. Biere, 2016.
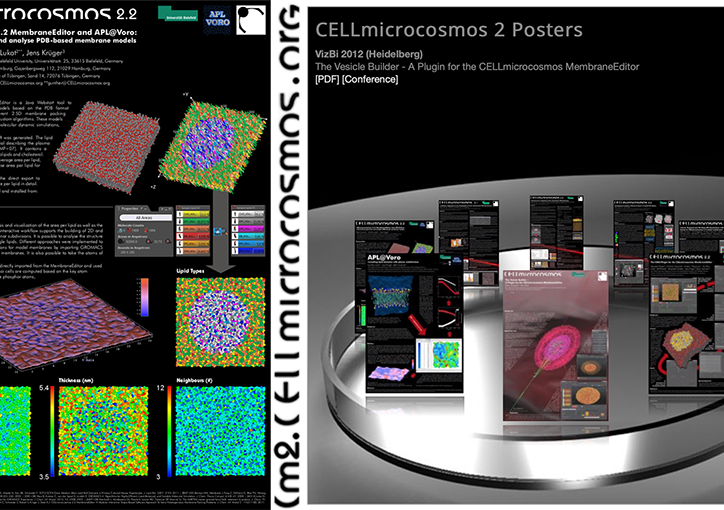
Yasara Prize for Best Poster at Faraday Discussion 169, Nottingham: 'CELLmicrocosmos 2.2.2 MembraneEditor and APL@Voro: interactively generate and analyse PDB-based membrane models', with G. Lukat, J. Krüger, 2014.
Best Use of Stereoscopy in a Technical Presentation at Stereoscopic Diplays and Applications XXV., San Francisco: 'Stereoscopic Cell Visualization: From Mesoscopic to Molecular Scale', with C. Bender, T. Hoppe, C. Gamroth, L. Jelonek, 2014.
Best Design Award at [Science Fair], Bielefeld: 'CE3: An eLearning Software for Cytological Education', with M. Civico, R. Orlik, 2011.
Current and recent projects
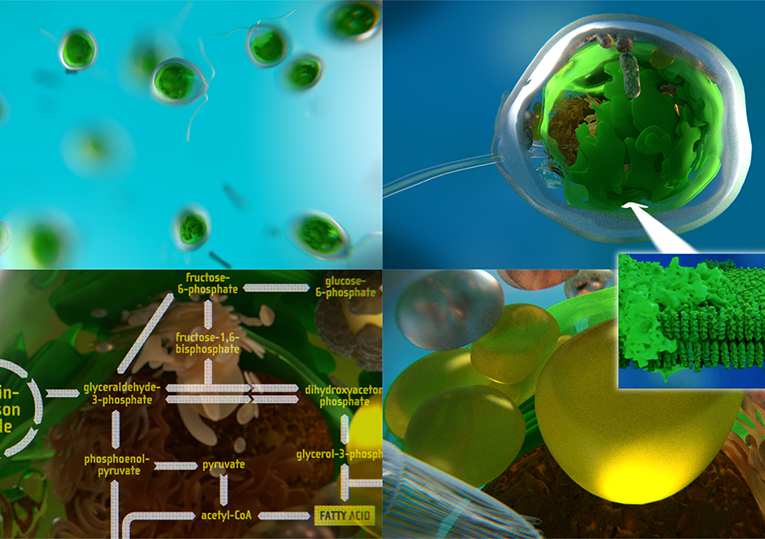
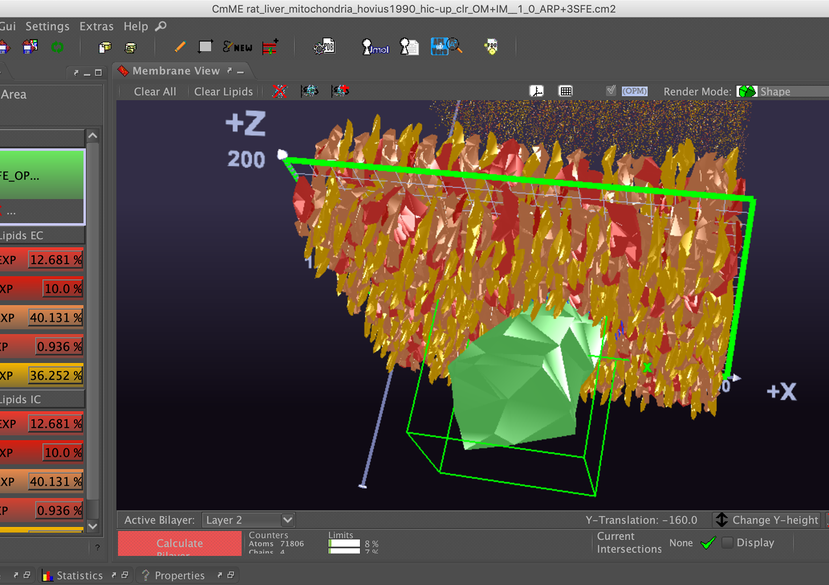
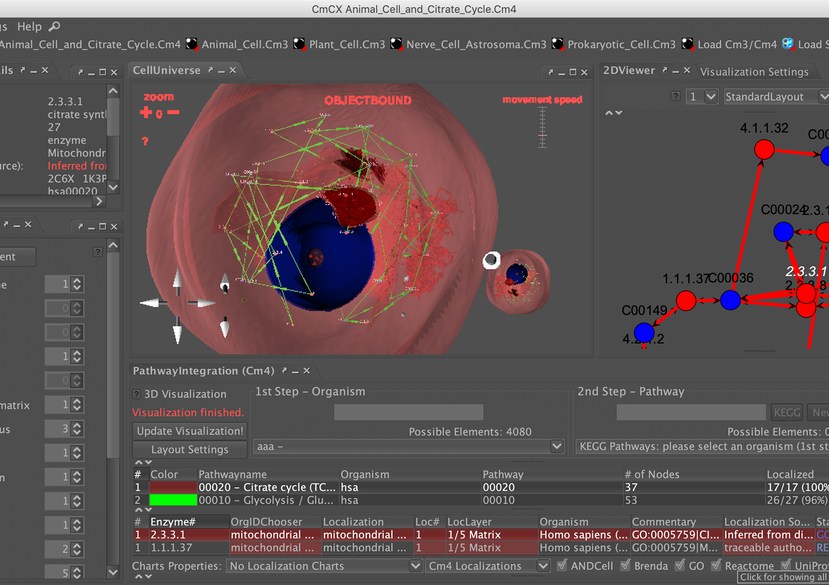
This project provides different approaches to model and visualise biology-inspired cell environments: For this purpose, software tools developed during the CELLmicrocosmos (Cm) project are used, as well as externally developed tools, for 3D animation, rendering and finishing, or molecular simulations. Also, it makes use of database-derived information. The project operates on the mesoscopic, molecular and functional level. Virtual Reality-related technology is used for visualising cellular and membrane structures. These different approaches were initially developed at Bielefeld University and then pushed forward at Monash University as well as the University of Konstanz.
Collective Behaviour Visualisation
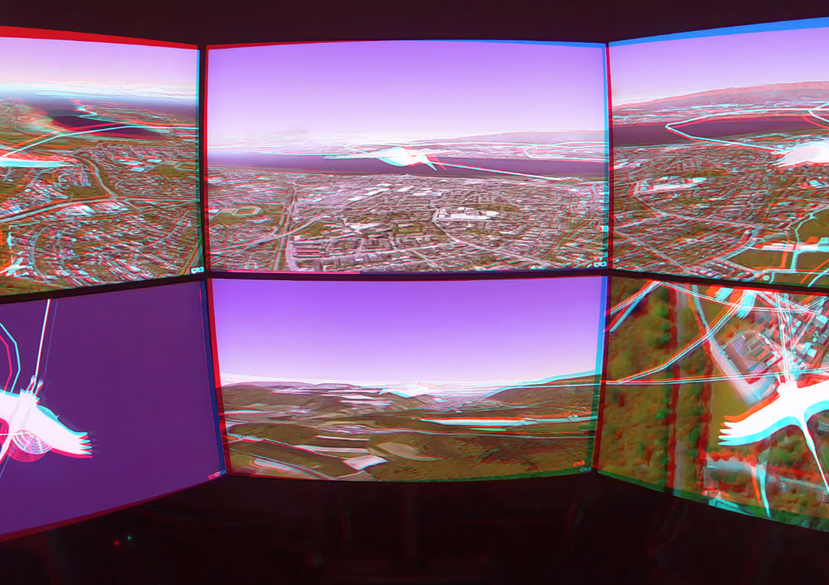
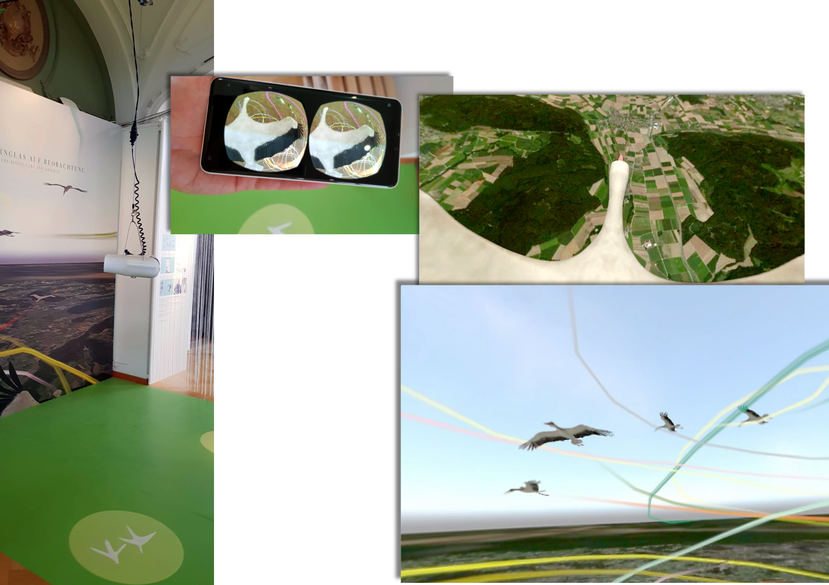
Together with the University of Konstanz and the Max Planck Institute of Animal Behaviour, different approaches were developed to analyse and visualise animal behaviour of birds and fish. Software tools as well as exhibition pieces using Virtual Reality technology were created for this purpose. A special focus was laid on Immersive Analytics techniques.
Publications, exhibitions, other outcomes
Sommer, B. et al. (2019) ‘Tiled Stereoscopic 3D Display Wall - Concept, Applications and Evaluation’, in Electronic Imaging, Stereoscopic Displays and Applications XXX. Stereoscopic Displays and Applications XXX, Burlingame, USA: Society for Imaging Science and Technology.
Sommer, B. et al. (2019) ‘BinocularsVR–A VR experience for the exhibition “From Lake Constance to Africa, a long distance travel with ICARUS”’, Electronic Imaging, 2019(2), pp. 177-1-177–8.
Woods, A. J. et al. (2019) ‘30th Annual Stereoscopic Displays and Applications Conference-Introduction’, Electronic Imaging, 2019(3), pp. B03-1-B03-7.
Klein, K., Sommer, B., et al. (2019) ‘Fly with the flock: immersive solutions for animal movement visualization and analytics’, Journal of the Royal Society Interface, 16(153), p. 20180794.
Zhu, Y. et al. (2018) ‘Genome-scale metabolic modeling of responses to polymyxins in Pseudomonas aeruginosa’, GigaScience, 7(4), p. giy021.
Wang, Y. et al. (2018) ‘Comparing Sequential and Temporal Patterns from Human Mobility Data for Next-Place Prediction’, in Proceddings of UMAP’18.
Wang, S. J., Sommer, B. et al. (2018) ‘The Virtual-Spine Platform—Acquiring, visualizing, and analyzing individual sitting behavior’, PLOS ONE, 13(6), p. e0195670.
Ghaffar, M., … and Sommer, B. (2018) ‘3D Modelling and Visualisation of Heterogeneous Cell Membranes in Blender’, in Proceedings of VINCI’18.
Biere, N., … and Sommer, B. (2018) ‘Heuristic Modeling and 3D Stereoscopic Visualization of a Chlamydomonas reinhardtii Cell’, Journal of Integrative Bioinformatics, 15(2).
Baltabayev, A., … and Sommer, B. (2018) ‘Virtual Reality for Sensor Data Visualization and Analysis’, in Proceedings of Engineering Reality of Virtual Reality 2018. Engineering Reality of Virtual Reality 2018 (Electronic Imaging 2018), Burlingame, USA.
Sommer, B. et al. (2018) ‘From Virtual Reality to Immersive Analytics in Bioinformatics’, Journal of Integrative Bioinformatics, 15(2).
Woods, A. J. et al. (2018) ‘Stereoscopic Displays and Applications XXIX-Introduction’, Electronic Imaging, 2018 (4), pp. 476-1-476–7.
Czauderna, T. et al. (2018) ‘Immersive Analytics Applications in Life and Health Sciences’, in Immersive Analytics. Springer, pp. 289–330.
Sommer, B. et al. (2017) ‘3D-Stereoscopic Immersive Analytics Projects at Monash University and University of Konstanz’, Electronic Imaging, 2017(5), pp. 179–187.
Hieu, N. et al. (2017) ‘Design Considerations for Immersive Analytics of Bird Movements Obtained by Miniaturised GPS Sensors’, in Proceedings of Eurographics Workshop on Visual Computing for Biology and Medicine (VCBM) 2017. Eurographics Workshop on Visual Computing for Biology and Medicine (VCBM) 2017, Bremen, pp. 89–96.
Sommer, B. and Schreiber, F. (2016) ‘Integration and virtual reality exploration of biomedical data with CmPI and VANTED’, it-Information Technology.
Sommer, B. et al. (2016) ‘Stereoscopic Space Map – Semi-immersive Configuration of 3D-stereoscopic Tours in Multi-display Environments’, Electronic Imaging, Proceedings of Stereoscopic Displays and Applications XXVII, 2016(5), pp. 1–9.
Schreiber, F. et al. (2016) ‘Specifications of standards in systems and synthetic biology: status and developments in 2016’, Journal of Integrative Bioinformatics, 13(3), pp. 1–7.
Qilin, L. et al. (2016) ‘DaTo: an atlas of biological databases and tools’, Journal of Integrative Bioinformatics, 13(4), p. 297.
Nim, H. T. et al. (2016) ‘Communicating the Effect of Human Behaviour on the Great Barrier Reef via Mixed Reality Visualisation’, in Big Data Visual Analytics (BDVA), 2016. Big Data Visual Analytics (BDVA), Sydney, Australia: IEEE, pp. 1–6.
Mueller, S. C. et al. (2016) ‘From single variants to protein cascades: Multi-scale modelling of SNV sets in genetic disorders’, Journal of Biological Chemistry, 291, pp. 1582–1590.
Kovanci, G., Ghaffar, M. and Sommer, B. (2016) ‘Web-based hybrid-dimensional Visualization and Exploration of Cytological Localization Scenarios’, Journal of Integrative Bioinformatics, 13(4), p. 298.
Sommer, B. et al. (2015) ‘Hybrid-Dimensional Visualization and Interaction - Integrating 2D and 3D Visualization with Semi-Immersive Navigation Techniques’, in Big Data Visual Analytics (BDVA), 2015. IEEE International Symposium on Big Data Visual Analytics, IEEE, pp. 1–8.
Schreiber, F. et al. (2015) ‘Specifications of Standards in Systems and Synthetic Biology’, Journal of Integrative Bioinformatics, 12(2), p. 258.
Waltemate, T., Sommer, B. and Botsch, M. (2014) ‘Membrane Mapping: Combining Mesoscopic and Molecular Cell Visualization’, in. Vienna, Austria: Eurographics Association (Proceedings of Eurographics Workshop on Visual Computing for Biology and Medicine), pp. 89–96. doi: 10.2312/vcbm.20141187.
Sommer, B. et al. (2014) ‘Stereoscopic cell visualization: from mesoscopic to molecular scale’, Electronic Imaging, Proceedings of Stereoscopic Displays and Applications XXVIII, 23(1), pp. 011007-1-011007–10. doi: 10.1117/1.JEI.23.1.011007.
Sommer, B. (2014) Proceedings of CELLmicrocosmos NeXt 2014; CELLmicrocosmos NeXt Workshop in Context of the" German Conference on Bioinformatics"; Bielefeld, 28.09. 2014. Universitätsbibliothek Bielefeld, Bielefeld eCollections (BieColl).
Sommer, B. (2014) ‘Network Analysis and Integration in a Virtual Cell Environment’, in Approaches in Integrative Bioinformatics. Springer, pp. 275–297.
Popik, O. V. et al. (2014) ‘Analysis of signaling networks distributed over intracellular compartments based on protein-protein interactions’, BMC Genomics, 15(Suppl 12), p. S7.
Sommer, B. (2013) ‘Membrane Packing Problems: A short Review on computational Membrane Modeling Methods and Tools’, Computational and Structural Biotechnology Journal, 5(6), p. e201302014.
Sommer, B. et al. (2013) ‘Subcellular Localization Charts: A new visual methodology for the semi-automatic localization of protein-related data sets’, Journal of Bioinformatics and Computational Biology, 11(1), p. 1340005.
Lukat, G., Krüger, J. and Sommer, B. (2013) ‘APL@Voro: A Voronoi-Based Membrane Analysis Tool for GROMACS Trajectories’, Journal of chemical information and modelling, 53(11), pp. 2908–2925.
Hofestädt, R. et al. (2013) ‘JIBtools: a Strategy to Reduce the Bioinformatics Analysis Gap’, Journal of Integrative Bioinformatics, 10(1), p. 226.
Sommer, B. (2012) CELLmicrocosmos - Integrative cell modeling at the molecular, mesoscopic and functional level. Doctorate Thesis. Bielefeld University.
Rubert, S., … and Sommer, B. (2012) ‘Grid Workflow Approach using the CELLmicrocosmos 2.2 MembraneEditor and UNICORE to commit and monitor GROMACS Jobs’, Proceedings of the Grid Workflow Workshop (GWW2011), Cologne, Germany, March 4, 2011. (CEUR-WS), 826.
Sommer, B. et al. (2011) ‘CELLmicrocosmos 2.2 MembraneEditor: a modular interactive shape-based software approach to solve heterogeneous Membrane Packing Problems’, Journal of Chemical Information and Modeling, 5(51), pp. 1165–1182.
Sommer, B. (2010) ‘Membranen für alle’, Nachrichten aus der Chemie, 58(10), pp. 1030–1032.
Sommer, B. et al. (2010) ‘Visualization and analysis of a Cardio Vascular Disease-and MUPP1-related biological network combining text mining and data warehouse approaches’, Journal of Integrative Bioinformatics, 7(1), p. 148.
Sommer, B. et al. (2010) ‘CELLmicrocosmos 4.1: an interactive approach to integrating spatially localized metabolic networks into a virtual 3D cell environment’, in Fred, A., Filipe, J., and Gamboa, H. (eds) BIOINFORMATICS 2010 – Proceedings of the 1st International Conference on Bioinformatics, part of the 3rd International Joint Conference on Biomedical Engineering Systems and Technologies (BIOSTEC 2010), pp. 90–95.
Sommer, B. (2009) ‘Cover Image and Feature Article for Informatik Spektrum’, Informatik Spektrum: Special Issue: Bioinformatik.
Exhibitions
link zur künstlichen intelligenz, Turm zur Katz Konstanz (2019), with students and lecturers of: HTWG Konstanz, University of Konstanz, Staatliche Hochschule für Musik Trossingen/Hochschule Furtwangen university; http://link-ki.de, Role: co supervision
BinocularsVR – A VR experience for the exhibition ‘rom Lake Constance to Africa, a long-distance travel with ICARUS, in the Deutschordensschloss at the island of Mainau in Germany (2018), with students and lecturers of University of Konstanz, Role: main co supervision and development.
External collaborations
- (2019) Co-organiser, Visualizing Biological Data – VizBi .
- (2018) Co-organiser, Big Data Visual Analytics – BDVA.
- Steering Committee Member, Integrative Bioinformatics – IB.
- (2018, 2019) Bio Visualisation Workshops on Cell and Membrane Visualisation at VizBi, IB, EuroVis MolVA
- Managing Editor, Journal of Integrative Bioinformatics: